+49 (0)8143-99172-0
Headquarter
+49 (0)8143-99172-22
Support

0x consumables
5x faster switching
10x faster cleaning
12x fluorescence channels
∞x statistics
…and much more!

We measure your products in a measuring range of 0.8 nm to 300 μm, if possible in original concentration
Particle Metrix GmbH, Inning, Germany
Institute for Transfusion Medicine (AG Giebel), University Hospital Essen,
University of Duisburg-Essen, Germany
Colocalization Nanoparticle Tracking Analysis (C-NTA) represents significant further development of fluorescence-NTA (F-NTA)
methods. In addition to measuring the size and concentration of fluorescently labelled particles, C-NTA offers the ability to
quickly detect two or more fluorescently labelled targets which coincide on a single extracellular vesicle. Multiple lasers and
the selection of corresponding fluorescence filters allow for quantification of individual fluorescent sub-populations as well as
particles which are double-positive for different surface markers.
After successfully showing C-NTA of double-stained MSC-EVs in our previous study (https://bit.ly/43GG0Yk) and confirming
the results with image flow cytometry, this note describes C-NTA of multi-stained MSC-EVs using a PMX-430 QUATT NTA
system with four lasers.
Keywords: colocalization, multi-staining, Nanoparticle Tracking Analysis (NTA), extracellular vesicles, CD41, CD9, CellMask Deep Red, antibody, fluorescence, membrane, NP-40
Extracellular vesicles (EVs) from selected cell types are currently being studied intensively because of their critical therapeutic potential. EVs can be classified
according to their origin, yet recent studies indicate a high degree of heterogeneity even within certain EV classes [1-6]. One primary goal in EV research is to establish and define methods which facilitate the robust detection of heterogeneity within purified samples of EVs, particularly in sample fractions containing EV sizes < 200 nm [1, 3]; however, progress in this context is still hampered by the fact that most currently available methods for analysing smaller EVs are either very time consuming or can only quantify a limited set of parameters [7, 8].
EVs were derived from MSC conditioned media grown in the presence of human platelet-lysate via a PEG precipitation procedure [see previous study 15, 16].
For antibody labelling, a cocktail of (i) 1 μl of MSC-EVs with a measured concentration of 4.2×1011 EV/ml, was mixed with (ii) 5 μl of a 1:25 dilution of anti-Hu CD9PB, a Pacific Blue-conjugated antibody (clone MEM-61), (iii) 5 μl of a 1:125 dilution of anti-Hu CD41-AF488 antibody (clone MEM-06), and (iv) 1 μl of a 1:250 dilution of CMDR Plasma Membrane Stain. Both pre-labeled antibodies were obtained from EXBIO, Vestec (Czech Republic), whereas, CMDR was obtained from Thermo Fisher Scientific (Schwerte, Germany). EVs and both antibodies were incubated for 1 hour in the dark at room temperature. After antibody incubation, CMDR was then added and further incubated for an additional hour, resulting in 2 hours incubation. The incubation cocktail was then diluted with PBS buffer to a final volume of 1 ml for subsequent NTA analyses.
Detergent controls were prepared with non-ionic detergent NP-40 Alternative (aka NP-40), from Calbiochem, Merck, Darmstadt (Germany), used to rupture the membrane of EVs. Following incubation, 12 μl of 0.5% NP-40 solution (diluted in particle-free PBS) was added to the labeled EV cocktail, resulting in a final concentration of 0.25% NP-40, and then incubated an additional 30 minutes at room temperature in the dark. The lysed cocktail was diluted into 1 ml of PBS for NTA analyses.
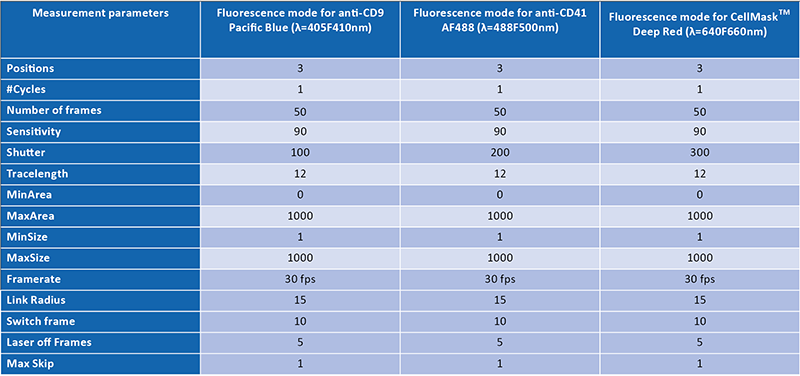
NTA measurements of size and concentration as well as colocalization measurements were made with a ZetaView® PMX-430 QUATT equipped with the Particle Metrix Software Suite and the ZetaNavigator software v1.1 (SOPs in Table 1). Measurements and the calculation of the colocalization ratios were performed as described in our previous study (see abstract link). The main difference in this study is that 3 pairs of fluorescence channels (using 405, 488, 640 sources) were used instead of one pair (488 & 640) for determining the colocalization ratios of the EVs.
Standard NTA measurements in scatter mode with the ZetaView® system resulted in a median diameter (X50) of 117 nm and a concentration of 2.5×1011 particles per ml.
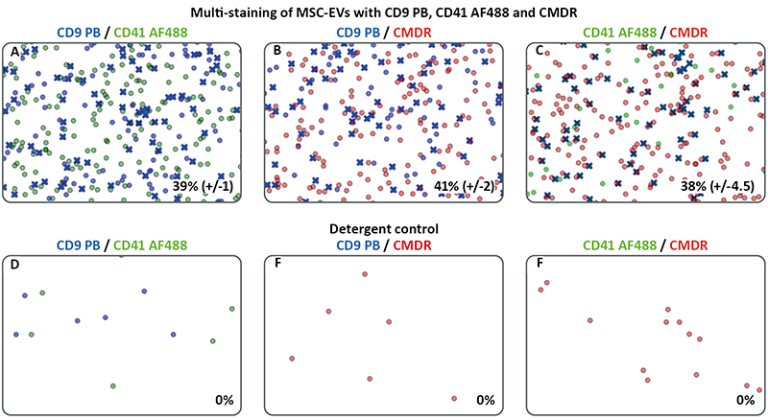
To specifically phenotype EVs in the sample, the particles were multi-labelled with anti-CD9 on Pacific Blue, anti-CD41 on AF488 and the membrane dye CellMask™ Deep Red. The specific choice of fluorochromes reflects our need to ensure that spill-over in the individual fluorescence channels would be avoided.
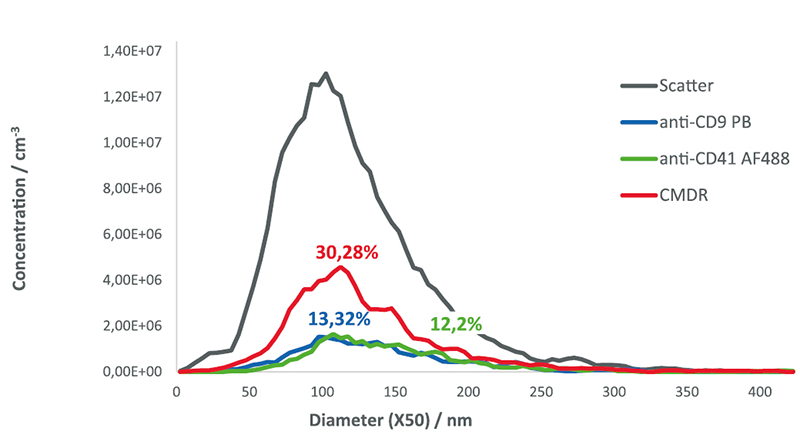
C-NTA measurements from all three labels yielded dual-positive detections of vesicles in all three pairings of fluorescence channels with the ZetaView® instrument.
CD9-positive EVs resulted in a median diameter (X50) of 128.8 nm and a measured concentration of 3.3×1010 EV/ml. In contrast, the mean diameter
of the CD41-positive EVs was 136.6 nm, which was slightly larger than the particles detected in the other fluorescence and scatter mode. The measured concentration of CD41-positive EVs was 3.05×1010 EV/ml. EVs positive for CMDR revealed a median size of 124.8 nm and a concentration of 7.57×1010 particles/ml. This concentration is more than twice as large as the CD9 & CD41 populations, suggesting the presence of additional EVs and/or other lipid membrane bearing particles without these tetraspanins.
The percentage of CD9- and CD41- and CMDR-positive EVs detected in the blue, green and red fluorescence channels were found to be 13.3%, 12.2% and 30.28%, respectively. Given that the sample was not purified and therefore likely contains different nanoparticles besides EVs, the CMDR results were expected, as this dye does not specifically stain EVs, rather, other lipid- or membrane-containing structures will also be counted in the 640 nm channel.
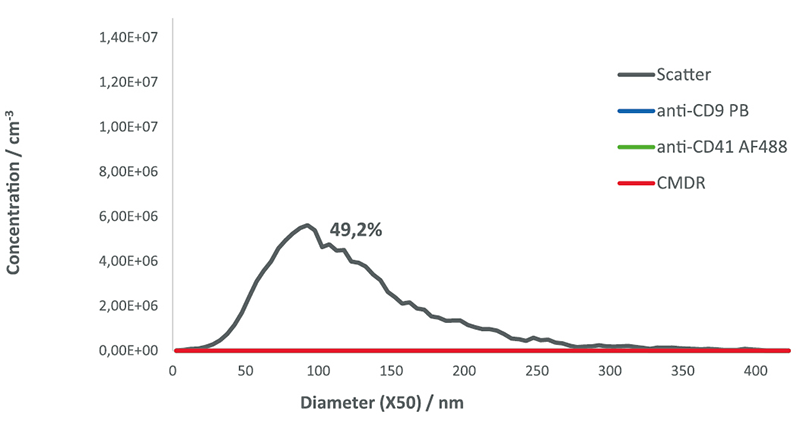
NP-40 detergent controls showed a particle number reduced by 50.8% in scatter mode and no particles in the fluorescence mode. The fact that around 50% of
the particles remain observable in the scatter mode indicates that half of the population in the sample contains non-membranous particles resistant to the detergent.
Using C-NTA, the same volume of fluorescently labelled particles is illuminated very quickly, one laser after the other with three different wavelengths, and the fluorescence signals are detected in series using the corresponding filters (see table 1). C-NTA of the multistained EVs showed colocalization ratios between 39%-41% (see figure 3 A-C) and revealed probably the highest specificity on the vesicles detected for CD9-CD41, because two paired antibodies were detected. In addition, the two pairs CD9-CMDR and CD41-CMDR show comparable colocalization ratio (41% (+/-2) and 38% (+/-4.5)). Notably, staining with CMDR is not as targeted as with use of conjugated antibodies, due ubiquitous binding to all lipid-membrane particles. In contrast to our previous study (see above), we here observe a lower colocalization ratio of CD9-CD41 (58.5% vs. 39%), most probably because a different batch of MSC-EVs was used.
As expected, no colocalization events were observed in the detergent controls using 0.5% NP-40 (see figure 3 D-F).
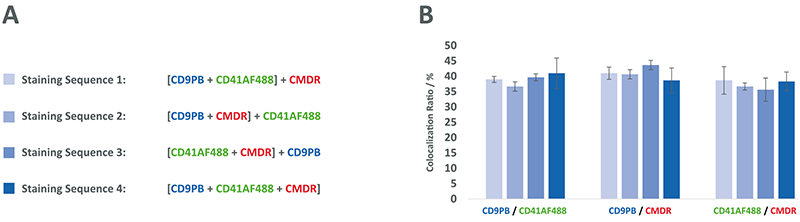
The more dyes or antibodies that are used for C-NTA, the more diverse the sequence in which the different dyes can be added. In order to investigate whether
colocalization ratios are dependent on the order in which the antibodies and mermbrane labels were added, we tested 4 different staining sequences on the MSC-EVs. The result is shown in figure 4 and shows that, at least with this mixture of labels, no significant difference in the colocalization results was observed for the different staining sequences.
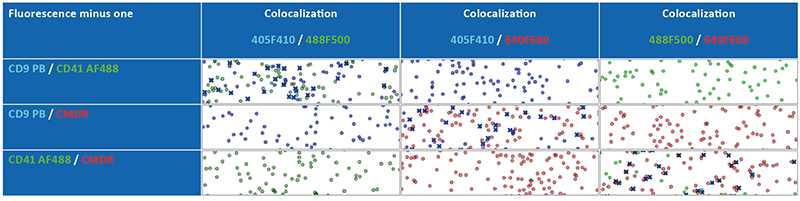
Fluorescence minus one (FMO) controls are important when identifying a positive from a negative population. When acquiring data, there might be fluorescence spread (spill-over), particularly with brighter fluorophores, which is noticeable after compensation (not required in this study) and cross-laser excitation. FMO controls have been done for all fluorophores in our panel when we started the new multicolor experiment. During staining, one dye or antibody was omitted, but colocalization was still tested in all available fluorescence channels (see figure 5). As a result, depending on the combination of the dyes, there was only one pair of fluorescence channels in which colocalization was observed.
In this study, we describe for the first time the characterization and phenotyping of multi-labeled EVs using C-NTA. We were able to quickly and reproducibly detect the ratio of CD9/CD41, CD9/CMDR and CD41/ CMDR double positive EVs using multiple lasers and flexibly selectable pairs of fluorescence filters. Since brownian motion is also tracked during each C-NTA measurement, accurate size distributions and concentration information of all fluorescent EV-subpopulations are available during a single measurement, simultaneously with the colocalization data.
False-positive detections and artefacts were prevented with the detergent controls. Furthermore, these controls verified that the colocalized particles were indeed EVs.
FMO controls did not show any fluorescence spread in adjacent channels, thus proving the specificity of both (i) the labeling, and (ii) the fluorophores with regard to the fluorescence channels used. The possible permutations for labelling sequences increase greatly with each additional label, thus we tested different staining sequences and found that the staining sequence has no significant influence on the colocalization ratios examined here.
As in our previous study, negative controls prepared without MSC-EVs, but containing all other components, show a significantly reduced number of particles compared to the positive samples, and the diameter of the particles remained roughly the same (data not shown).
The use of highly targeted C-NTA experiments shows that combinations of antibody and membrane staining can determine the amount of EVs of interest. In addition, C-NTA has proven to be an accurate tool for the detection and identification of biomarkers at the single particle level within a single measurement, including the benefit of accurate sizing and concentration measurement.
C-NTA therefore offers a highly targeted approach to helping unravel the remaining mysteries in the exciting world of EVs.
1. Koliha N, Heider U, Ozimkowski T, et al.:
Melanoma affects the composition of blood cell-derived extracellular vesicles [original research]. Front Immunol.
2016 Jul;7(282). English.
DOI:10.3389/fimmu.2016.00282
2. Koliha N, Wiencek Y, Heider U, et al.:
A novel multiplex bead-based platform highlights the diversity of extracellular vesicles. J Extracell Vesicles.
2016;016;5:29975. PubMed PMID: 26901056; PubMed
Central PMCID: PMCPMC4762227.
3. Wiklander OP, Nordin JZ, O’Loughlin A, et al.:
Extracellular vesicle in vivo bio-distribution is determined
by cell source, route of administration and targeting.
J Extracell Vesicles. 2015;4:26316. PubMed PMID:
25899407; PubMed
Central PMCID: PMCPMC4405624.
4. Wiklander OPB, Bostancioglu RB, Welsh JA, et al.:
Systematic methodological eva-luation of a multiplex
bead-based flow cytometry assay for detection of
extra-cellular vesicle surface signatures
Front Immunol. 2018 Jun;9(1326).
English. doi: 10.3389/fimmu.2018.01326.
5. Kowal J, Arras G, Colombo M, et al.:
Proteomic comparison defines novel markers to characterize
heterogeneous populations of extracellular vesicle
sub-types. Proc Natl Acad Sci U S A. 2016 Feb 8. PubMed
PMID: 26858453.
DOI:10.1073/pnas.1521230113.
6. Giebel B.
On the function and hetero-geneity of extracellular
vesicles. Ann Transl Med. 2017 Mar;5(6):150.
PubMed PMID: 28462230; PubMed Central PMCID:
PMCPMC5395490.
7. Witwer KW, Buzas EI, Bemis LT, et al.:
Standardization of sample collection, isolation and
analysis methods in extracellular vesicle research. J Extracell
Vesicles. 2013;2. PubMed PMID: 24009894; PubMed
Central PMCID: PMCPMC3760646.
DOI:10.3402/jev.v2i0.20360
8. Sharma S, Rasool HI, Palanisamy V, et al.:
Structuralmechanical characterization of nanoparticle
exosomes in human saliva, using correlative AFM, FESEM,
and force spectroscopy. ACS Nano. 2010;4(4):1921–1926
Disclaimer: While we have taken every care to ensure that the information in this document is correct, nothing herein should be construed as implying any representation or warranty as to its accuracy, correctness or completeness of this information and we shall not be held liable for any errors therein or for any damages in connection with the use of this material. Particle Metrix reserves the right to change the content of this material at any time without notice.
Copyright: © May 2023 Particle Metrix. This publication or parts thereof may not be copied or distributed without our express written permission.
Headquarter
Support
Opt-out abgeschlossen; Ihre Besuche auf dieser Webseite werden nicht vom Webanalysetool erfasst. Bitte beachten Sie, dass auch der Matomo-Deaktivierungs-Cookie dieser Webseite gelöscht wird, wenn Sie die in Ihrem Browser abgelegten Cookies entfernen. Außerdem müssen Sie, wenn Sie einen anderen Computer oder einen anderen Webbrowser verwenden, die Deaktivierungsprozedur nochmals absolvieren.
Sie haben die Möglichkeit zu verhindern, dass von Ihnen hier getätigte Aktionen analysiert und verknüpft werden. Dies wird Ihre Privatsphäre schützen, aber wird auch den Besitzer daran hindern, aus Ihren Aktionen zu lernen und die Bedienbarkeit für Sie und andere Benutzer zu verbessern.
Die Tracking opt-out Funktion benötigt aktivierte Cookies.